Preparing for Indexing: Faster, Safer, More Useful Data at Rest
Vincent Brandon, Data Coordinator
June 2, 2021

What is indexing?
Data does not just sit. It takes up space, has a place, and finding it can be terribly hard. Imagine a library without ISBN codes where we haphazardly throw books around. Indexing is all about finding things or things about things fast. If I know a book is in a particular aisle, that limits how much searching I need to do. We all rely on indexes every time we open a database, look at email, or go to the library. The UDRC, though, has some interesting issues.
Our books need to be secure. It is tempting to say ‘Adam’ is in the ‘College Row’ under ‘2010 Graduates.’ That would make finding Adam’s record much easier but not at all secure for his personally identifiable information.
Embeddings as Index - Indexing for Probabilistic Search
How can the UDRC secure data at rest while allowing blazing fast access times for our matching algorithms? Neural networks provide an interesting answer, but the indexes take on a new form. Multi-dimensional vector spaces, big trees that stretch over our data, and tell us how to look for things. You cannot climb these branches with simple strings, though. We need to prepare the data, and in that, we have an opportunity to secure further what we store.
Let us illustrate the process with a common problem: Names. Matching names is a horrible procedure. There are misspellings, acronyms, nicknames. And we cannot map a random string sequence onto our tree. We need to embed the name in a set structure. In practice, we use neural networks, often transformers, for this. But here is a simpler approach:
- We have two people who have lots of nicknames, Sam and Dave.
- We need to create a fast lookup wherever any of their names appear, replacing their names with an embedding method.
- For this example, as in Table 1, we will do an alphabet embedding with a fixed length of 5.
- Each letter is replaced by its numerical position in the English alphabet.
| Name | Embedding |
|---|---|
| Sammy | <19, 1, 13, 13, 25> |
| San | <19, 1, 12, 0, 0> |
| Samuel | <19, 1, 13, 21, 5> |
| Dave | <4, 1, 22, 5, 0> |
| Dabid | <4, 1, 2, 9, 4> |
Visualizing Vector Space
You might be able to see right away how different these names are numerically. At the UDRC, we do not require exact name matches, nor do we match on the name alone. Plenty of people have the same name even when everyone’s spelling is top-notch, and no one ever uses nicknames. What we want to know is who it ‘could’ be. Who is this person similar to? If we graph the vector’s first three points, as in Figure 1, we can immediately see how clusters of names might begin to form.
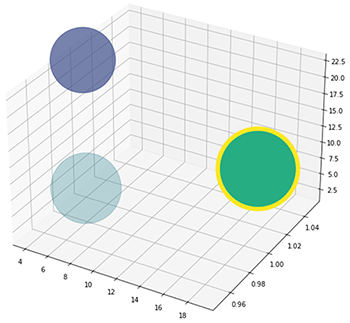
With the tens of thousands of names across the hundreds of dimensions that our embedding networks create, visualization is a bit tricky. The true power of vector embedding is not in visualization directly but in optimized computation. We can now calculate distances between the names very fast. Remember Pythagorean’s Theorem? We can calculate Euclidean distances and ask, how far is Sammy to Dave?
| Comparison | Distance |
|---|---|
| Sammy to Samuel | 8.0 |
| Sammy to Dave | 19.2 |
Sammy and Samuel may be the same person, but Sammy and Dave are probably not. With smarter embedding and optimized distance metrics, we can cluster pretty accurately or search similarity very fast.
The UDRC uses personally identifiable information (PII) to match data to a random master person index (MPI), and then we have no use for it except to upgrade our datasets. The data we provide to researchers has all the PII removed, replaced with an MPI. These de-identified tables are the end product but are not static.
As we update the matching algorithm and adjust our pool of MPIs, we have to reconstruct the de-identified tables. This adjustment means keeping copies of the source data, and all that personally identifiable information is secured and available for reprocessing.
Embedding data into a searchable, non-reversible structure lets us rebuild faster, introspect our data, and provide extra security to our partners. Machine Learning is everywhere.